talks, outreach
EpiFor @ NetSci2011 June 20, 2011
 EpiFor team participated to NetSci2011, the International Conference on Network Science that took place at the Central European University in Budapest, HU, June 6-10.
EpiFor team participated to NetSci2011, the International Conference on Network Science that took place at the Central European University in Budapest, HU, June 6-10.
Bringing together leading researchers, practitioners, and teachers in network science (including analysts, modeling experts, visualization specialists, and others), every year NetSci fosters interdisciplinary communication and collaboration. The conference focuses on novel directions in networks research within the biological and environmental sciences, computer and information sciences, social sciences, finance and business.
EpiFor contributed to the conference with a poster and three contributed talks:
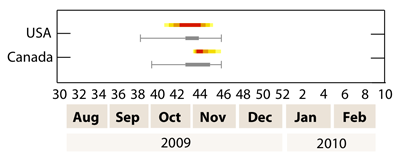 The poster titled "GLEaM, a global stochastic model for influenza epidemics: its application to the A/H1N1 pandemic" presented by Michele Tizzoni focused on the assessment of the GLEaM model predictions through the comparison between the simulated scenarios and the observed pandemic activity during the 2009-2010 fall/winter wave. The good agreement between simulations and surveillance data collected by monitoring systems in several countries confirms the valuable role of large computational epidemic models in pandemic preparedness and management.
The poster titled "GLEaM, a global stochastic model for influenza epidemics: its application to the A/H1N1 pandemic" presented by Michele Tizzoni focused on the assessment of the GLEaM model predictions through the comparison between the simulated scenarios and the observed pandemic activity during the 2009-2010 fall/winter wave. The good agreement between simulations and surveillance data collected by monitoring systems in several countries confirms the valuable role of large computational epidemic models in pandemic preparedness and management.
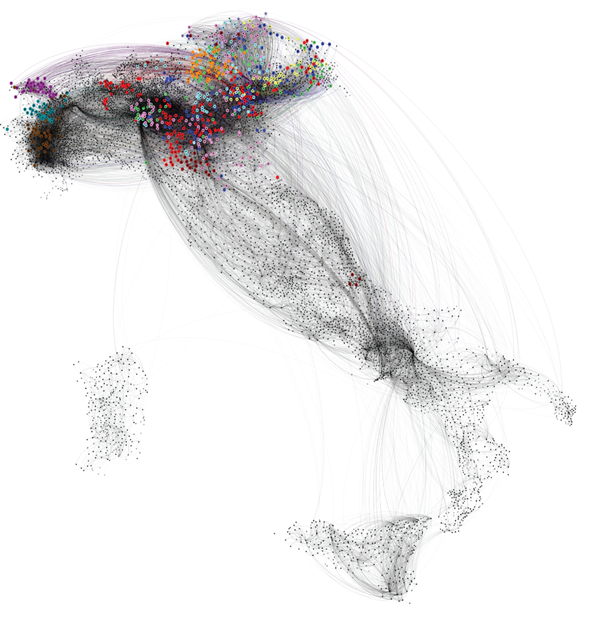
The contributed talk "A longitudinal complex networks analysis of the dynamical patterns of cattle trade movements" presented by Paolo Bajardi was aimed at the characterization of the dynamical phenomena of livestock movements developing a general framework to assess the impact of the temporal patterns on an epidemic spreading on the system. The results open the path to novel approaches that fully integrate the dynamical nature of the network to realistically describe a spreading process and devise specific interventions.
 In the contributed talk "The GLEaMviz computational tool, a publicly available software to explore realistic epidemic spreading scenarios at the global scale", Corrado Gioannini presented the GLEaMviz simulator software, a publicly downloadable tool (http://www.gleamviz.org/simulator/) –implementing the Global Epidemic and Mobility model– that can be used to perform realistic simulations of human-to-human disease spreading at the global scale.
In the contributed talk "The GLEaMviz computational tool, a publicly available software to explore realistic epidemic spreading scenarios at the global scale", Corrado Gioannini presented the GLEaMviz simulator software, a publicly downloadable tool (http://www.gleamviz.org/simulator/) –implementing the Global Epidemic and Mobility model– that can be used to perform realistic simulations of human-to-human disease spreading at the global scale.
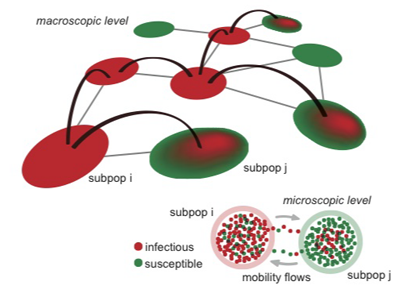 The contributed talk "Human travel and time spent at destination: impact on the epidemic invasion dynamics" by Chiara Poletto, focused on the impact of a fundamental aspect of human mobility, namely trip duration, on the geographical spread of infections. A modeling framework is introduced in order to provide understandings on how length of stay and its heterogeneities impact the ability of a local outbreak to reach pandemic proportions.
The contributed talk "Human travel and time spent at destination: impact on the epidemic invasion dynamics" by Chiara Poletto, focused on the impact of a fundamental aspect of human mobility, namely trip duration, on the geographical spread of infections. A modeling framework is introduced in order to provide understandings on how length of stay and its heterogeneities impact the ability of a local outbreak to reach pandemic proportions.
Posts by category: awards/honors, news, outreach, publications, research, talks, team news



